|
 |
- Search
| J Audiol Otol > Volume 16(1); 2012 > Article |
Abstract
The main objective of this review is to describe the new sequencing technologies called next generation sequencing (NGS) and its utility as a molecular genetic diagnosis tool in a medical field. Sanger method has dominated the genome sequencing industry for the past 30 years since its invention in 1975. It produced first human genome and still remains the gold standard for genome sequencing. However, it cannot meet the needs for enormous genetic data gathering and process because of its relatively long sequencing time and high cost per sample. NGS which parallelise the sequencing process, thereby increasing processing speed at a reduced cost per sample emerged to compensate for the weakness of the previous method. Currently NGS is used in some medical areas and its use is being widened. NGS also plays an important role in a study of genetically heterogenous hearing diseases. NGS is expected to mark a significant milestone in genomic research filed in a near future.
Since DNA was discovered as the code to all biological life on Earth, genome sequencing was regarded as the key to the management of all genetic diseases. Although nowadays genome sequencing is not considered as miraculous as it was in the past, the efforts to reveal the genome sequence helped developing many sequencing methods and the fast moving field of genomic medicine has been impacting on clinical care.
Since Sanger1) introduced his method for DNA sequencing in 1975, Sanger method and its variants have dominated the genome sequencing field for the past 30 years.2) The original Sanger method uses specifically labeled nucleotides to read through a DNA template during DNA synthesis. Sanger method remains the gold standard for sequencing accuracy. The current generation of automated high throughput Sanger sequencing machines read -2 million bases of sequence per day and were used to sequence the first human genome.3,4) However, this throughput is very low compared with the size of the human genome (one strand of the human genome is 3,000,000,000 base pairs). Therefore, new sequencing technologies called next generation sequencing (NGS) emerged.
NGS technologies parallelise the sequencing process, thereby massively increasing throughput (up to w50 billion bases per day) at a reduced cost per base. The Sanger method is considered as a 'first-generation' technology, and these newer methods are referred to as next-generation sequencing.
Currently, five major NGS platforms are commercially available. the Roche GS-FLX 454 Genome Sequencer (originally 454 sequencing), the Illumina Genome Analyzer (originally Solexa technology), the ABI SOLiD analyzer, Polonator G.007, the Helicos HeliScope platforms. A general process of NGS is outlined in Fig. 1.5)
Sequencing technologies include a number of methods that are grouped broadly as template preparation, sequencing and imaging, and data analysis. The unique combination of specific protocols distinguishes one technology from another and determines the type of data produced from each platform.
There are two methods used in preparing templates for NGS reactions: clonally amplified templates originating from single DNA molecules, and single DNA molecule templates.
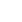
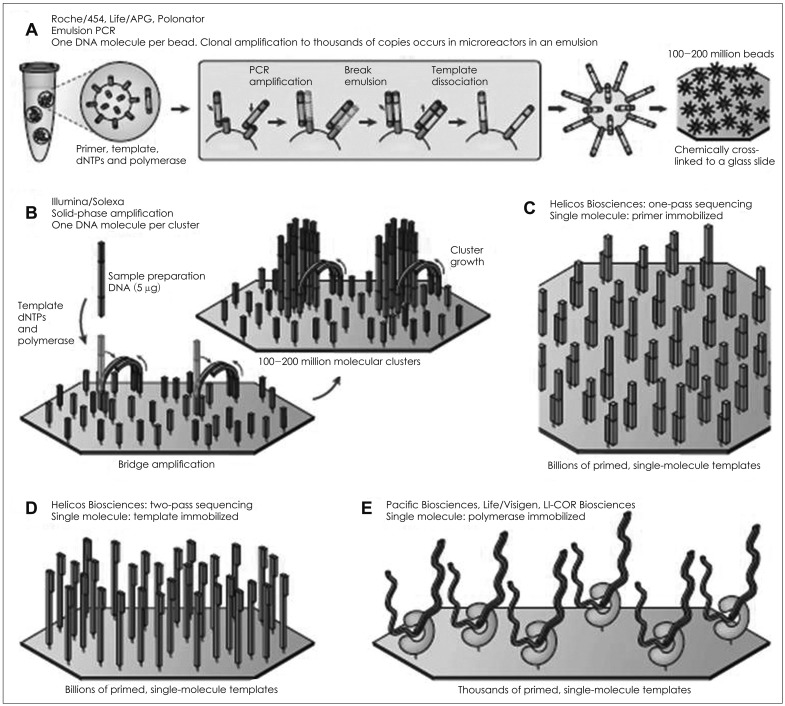
Clonally amplified templates are created through emulsion PCR (emPCR) and solid-phase amplification. Template preparation using these two methods are described below (Fig. 2A, B).6) After the successful amplification and enrichment of emPCR beads, millions can be immobilized in a polyacrylamide gel on a standard microscope slide (Polonator), chemically crosslinked to an aminocoated glass surface (Life/APG; Polonator) or deposited into individual PicoTiterPlate wells (Roche/454) in which the NGS chemistry can be performed. Solid-phase amplification can produce 100-200 million spatially separated template clusters (Illumina/Solexa), providing free ends to which a universal sequencing primer can be hybridized to initiate the NGS reaction.
NGS can be used with shortened time and reduced cost in all fields where automated sanger method has been used. It can be applied to whole genome sequencing (WGS), exome sequencing and target gene sequencing.
While NGS per base sequencing costs are relatively low, the cost of sequencing entire human genomes remains high and WGS is unnecessary for many diagnostic and research applications. It is estimated that the protein-coding regions of genes (exons) constitute approximately 1% of the human genome but harbour 85% of the mutations with large effects on disease.7) The exome is the part of the genome formed by exons, coding portions of genes in the genome that are expressed. Therefore, targeted sequencing of the exome or genes of interest seems a more efficient approach than WGS.
NGS is already used in some medical fields and its use is being widened. The application of NGS technology has produced a transformation in cancer genomics, generating large data sets that can be analyzed in different ways to answer a multitude of questions about the genomic alterations associated with the disease. Analytical approaches can discover focused mutations such as substitutions and small insertion/deletions, large structural alterations and copy number events.8)
Genome sequencing is already available for diagnostic use by cardiovascular clinicians, primarily for familial conditions. NGS is considered in the diagnosis of mendelian diseases such as Long QT syndrome and HCM, in molecular autopsy following sudden death and prenatal diagnosis.5) Both the Human Genetics Commission9) and the Foundation for Genomics and Population Health10) identified inherited cardiac conditions as a key area to help focus the development of an overall NHS strategy for NGS-based research and diagnostics.
As explained above, NGS is expected as a molecular genetic diagnosis tool in many medical fields currently and in the near future.
Deafness is a genetically heterogenous disease group, with 70 nuclear genes identified thus far for non-syndromic sensorineural hearing impairment and many more for syndromes including hearing loss.11) Most early-onset hearing loss is known to be genetic.12) Until recently, most genetic diagnosis for the deaf is limited to the most common mutations. Today, more target genes sequencing is being made available with the advent of NGS technologies.13) Gene capture combined with NGS has allowed a much greater number of samples to be examined with the advantages of 1) significant cost saving, 2) higher sequencing accuracy because of deeper achievable coverage, 3) a significantly shorter turnaround time, and 4) a more feasible data set for a bioinformatics analysis outcome that is functionally interpretable.14) Many new genes for non-syndromic hearing diseases15,16) and syndromic hearing diseases17-19) were discovered by the targeted NGS approach. For example, SMPX is known to encode the small muscle protein, X-linked (SMPX). However, one study using NGS identified SMPX as a gene in which variation is associated with X-linked deafness and illustrates that NGS is instrumental in the efficient identification of disease-causing variants in unexpected genes.20) Similarly, C9orf75, Encoding Taperin, was identified as the mutated gene in non-syndromic deafness DFNB79 by targeted capture and NGS.
Most newborn hearing loss cases are caused by genetic mutations, especially in developed countries.21) Therefore, a molecular genetic screening of a panel of genes involved in deafness is one of the potential role of NGS.
Contrary to complicated genetic traits observed in cardiomyopathy and cancer genomics, most genetic hearing loss cases are caused by monogenic lesion.22) NGS could be used in the diagnosis of genetic deafness. Although NGS is still not a definitive diagnostic tool, it could be used to make the additional tests, which might be expensive, unnecessary.
According to CDC data (http://www.cdc.gov/ncbddd/ehdi/data.htm), hearing loss prevalence increases from 1% at birth to 6% for school-age children. A potential patients with a carrier can be counseled and managed in advance with the help of NGS technologies.
Management of genetic hearing loss using the genetic level correction could be available in the future with the the widespread use of NGS technologies in the molecular genetic diagnosis.
Currently, NGS testing of up to 84 human genes implicated in both syndromic and non-syndromic hearing loss is offered on the market (http://www.healthcare.uiowa.edu/labs/morl/index_CDS.htm and www.otogenetics.com),14) and in practice, patients with A1555G mitochondrial mutations can be warned about their increased susceptibility for aminoglycoside ototoxicity.
According to Choi, et al.23) and Park, et al.,24) GJB2 and SLC26A4 are responsible for 10-15% and 20% of total deafness in Koreans respectively, while about 2/3 of deaf patients have not been molecular genetically diagnosed in Koreans. There might be completely different mutation spectrums depending upon ethnicities in Korea.
In Seoul National University Hospital, Choi, et al. performed pilot study of the target gene sequencing for 80 deafness genes among Korean deafness families. Cases were recruited from Sep. 2010 through April 2011, which included 10 autosomal dominant families and 1 autosomal recessive families. NimbleGen SeCap EZ Human Exome Library v2.0 capture kit was used for target exome capture and HiSeq2000 for sequencing. Analysis of bioinformatical data demonstrated very promising results in autosomal dominant inheritance families, which clarified the genetic etiology in 6 of 10 families and narrowed down to 2 potentially pathogenic variants in 3 of remaining 4 families. Whole process including chip design took 8 weeks and the cost was 400000 won per sample, while the cost for Sanger sequencing of the 80 genes required 5000000 won per sample. Authors nearly made a molecular genetic diagnosis from nine (90%) of ten AD families and again confirmed a heterogeneity with a reduced cost and shortened period compared with conventional sequencing method.
Genetic testing plays an important role in the diagnosis and management of genetically inherited disease. In the short term, Sanger sequencing method will continue to be used as a complementary strategy to sequence genomic regions that are difficult to target with NGS and also to validate NGS findings. As the accuracy and efficiency of targeted sequencing improves, NGS is expected increasingly be used as a standard technique.
Currently, NGS is applied into target gene sequencing in a research field and is introduced in the screening and counseling of genetic hearing diseases. As NGS costs fall, further move to exome sequencing and whole genome sequencing can be accomplished to investigate the genetic etiology of deafness.
In genomics, the personal genome era made available by NGS technologies will mark a significant milestone in entire genomic research field in the foreseeable future.
References
1. Sanger F. The Croonian Lecture, 1975. Nucleotide sequences in DNA. Proc R Soc Lond B Biol Sci 1975;191:317–333. PMID: 2920.


2. Hutchison CA 3rd. DNA sequencing: bench to bedside and beyond. Nucleic Acids Res 2007;35:6227–6237. PMID: 17855400.



3. Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, et al. The sequence of the human genome. Science 2001;291:1304–1351. PMID: 11181995.


4. McPherson JD, Marra M, Hillier L, Waterston RH, Chinwalla A, Wallis J, et al. A physical map of the human genome. Nature 2001;409:934–941. PMID: 11237014.


5. Ware JS, Roberts AM, Cook SA. Next generation sequencing for clinical diagnostics and personalised medicine: implications for the next generation cardiologist. Heart 2012;98:276–281. PMID: 22128206.


6. Metzker ML. Sequencing technologies - the next generation. Nat Rev Genet 2010;11:31–46. PMID: 19997069.


7. Choi M, Scholl UI, Ji W, Liu T, Tikhonova IR, Zumbo P, et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci U S A 2009;106:19096–19101. PMID: 19861545.



8. Ding L, Wendl MC, Koboldt DC, Mardis ER. Analysis of next-generation genomic data in cancer: accomplishments and challenges. Hum Mol Genet 2010;19:R188–R196. PMID: 20843826.



9. Human Genetics Commission. Information Gathering Session on the Strategic Priorities for Genetics Research. London: Human Genetics Commission;2007.
10. Burton H, Alberg C, Stewart A. Heart to Heart: Inherited Cardiovascular Conditions Services - a Needs Assessment and Service Review. Cambridge: The PHG Foundation;2009.
11. Van Camp GSR. Hereditary Hearing Loss Homepage 2012 Available from: URL: http://hereditaryhearingloss.org/.
12. Nance WE. The genetics of deafness. Ment Retard Dev Disabil Res Rev 2003;9:109–119. PMID: 12784229.


13. Brownstein Z, Friedman LM, Shahin H, Oron-Karni V, Kol N, Rayyan AA, et al. Targeted genomic capture and massively parallel sequencing to identify genes for hereditary hearing loss in middle eastern families. Genome Biol 2011;12:R89PMID: 21917145.



14. Lin X, Tang W, Ahmad S, Lu J, Colby CC, Zhu J, et al. Applications of targeted gene capture and next-generation sequencing technologies in studies of human deafness and other genetic disabilities. Hear Res 2012;In press.



15. Rehman AU, Morell RJ, Belyantseva IA, Khan SY, Boger ET, Shahzad M, et al. Targeted capture and next-generation sequencing identifies C9orf75, encoding taperin, as the mutated gene in nonsyndromic deafness DFNB79. Am J Hum Genet 2010;86:378–388. PMID: 20170899.



16. Walsh T, Shahin H, Elkan-Miller T, Lee MK, Thornton AM, Roeb W, et al. Whole exome sequencing and homozygosity mapping identify mutation in the cell polarity protein GPSM2 as the cause of nonsyndromic hearing loss DFNB82. Am J Hum Genet 2010;87:90–94. PMID: 20602914.



17. Shearer AE, DeLuca AP, Hildebrand MS, Taylor KR, Gurrola J 2nd, Scherer S, et al. Comprehensive genetic testing for hereditary hearing loss using massively parallel sequencing. Proc Natl Acad Sci U S A 2010;107:21104–21109. PMID: 21078986.



18. Pierce SB, Walsh T, Chisholm KM, Lee MK, Thornton AM, Fiumara A, et al. Mutations in the DBP-deficiency protein HSD17B4 cause ovarian dysgenesis, hearing loss, and ataxia of Perrault Syndrome. Am J Hum Genet 2010;87:282–288. PMID: 20673864.



19. Zheng J, Miller KK, Yang T, Hildebrand MS, Shearer AE, DeLuca AP, et al. Carcinoembryonic antigen-related cell adhesion molecule 16 interacts with alpha-tectorin and is mutated in autosomal dominant hearing loss (DFNA4). Proc Natl Acad Sci U S A 2011;108:4218–4223. PMID: 21368133.



20. Schraders M, Haas SA, Weegerink NJ, Oostrik J, Hu H, Hoefsloot LH, et al. Next-generation sequencing identifies mutations of SMPX, which encodes the small muscle protein, X-linked, as a cause of progressive hearing impairment. Am J Hum Genet 2011;88:628–634. PMID: 21549342.



21. Smith RJ, Bale JF Jr, White KR. Sensorineural hearing loss in children. Lancet 2005;365:879–890. PMID: 15752533.


22. Hilgert N, Smith RJ, Van Camp G. Forty-six genes causing nonsyndromic hearing impairment: which ones should be analyzed in DNA diagnostics? Mutat Res 2009;681:189–196. PMID: 18804553.



23. Choi BY, Stewart AK, Nishimura KK, Cha WJ, Seong MW, Park SS, et al. Efficient molecular genetic diagnosis of enlarged vestibular aqueducts in East Asians. Genet Test Mol Biomarkers 2009;13:679–687. PMID: 19645628.



24. Park HJ, Lee SJ, Jin HS, Lee JO, Go SH, Jang HS, et al. Genetic basis of hearing loss associated with enlarged vestibular aqueducts in Koreans. Clin Genet 2005;67:160–165. PMID: 15679828.


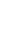
Fig. 1
Next generation sequencing (NGS) workflow (adapted from Ware JS, 2012). A: Genomic DNA is fragmented and platform specific 'adaptors' are attached. The DNA is then either attached to a bead or directly to the sequencing slide. In either case, the DNA is clonally amplified in this location to provide a cluster of molecules with identical sequences. If beads are used they are then immobilised on a sequencing slide. Different NGS platforms employ different sequencing chemistries. B: One approach to sequencing by synthesis, as employed by the Genome Analyser system (Illumina). The sequence of each fragment is read by decoding the sequence of fluorophores imaged at each physical position on a sequencing slide. Advanced optics allow for massively parallel sequencing. C: Each DNA molecule yields one or two sequence fragments depending on whether it is sequenced from one or both ends. These sequence fragments are computationally aligned with a reference sequence and mismatches identified.
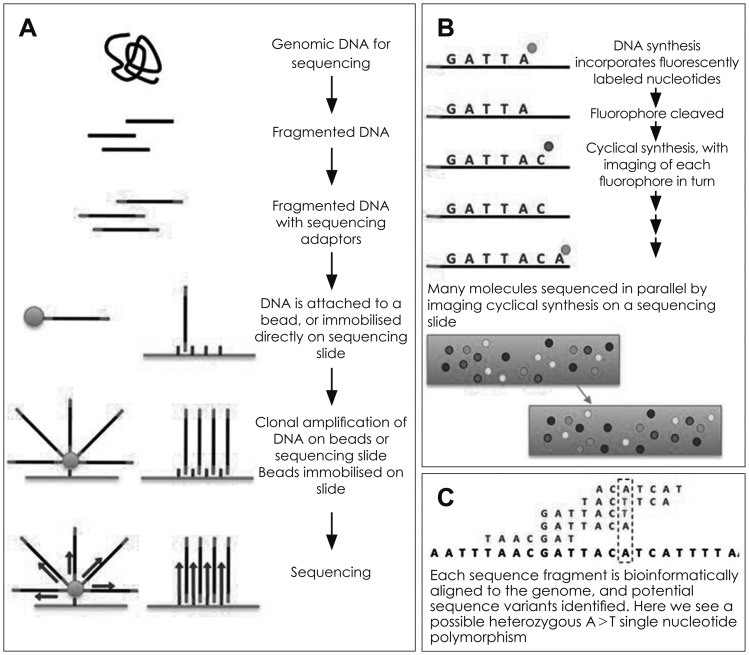
Fig. 2
Template immobilization strategies (adapted from Metzker ML, 2010). In emulsion PCR (emPCR)(A), a reaction mixture consisting of an oil-aqueous emulsion is created to encapsulate bead-DNA complexes into single aqueous droplets. PCR amplification is performed within these droplets to create beads containing several thousand copies of the same template sequence. EmPCR beads can be chemically attached to a glass slide or deposited into PicoTiterPlate wells. Solid-phase amplification (B) is composed of two basic steps: initial priming and extending of the single-stranded, single-molecule template, and bridge amplification of the immobilized template with immediately adjacent primers to form clusters. Three approaches are shown for immobilizing single-molecule templates to a solid support: immobilization by a primer (C), immobilization by a template (D), and immobilization of a polymerase (E). dNTP: 2'-deoxyribonucleoside triphosphate.